The JEFworks Lab at Johns Hopkins University seeks to better understand how cell identity is molecularly encoded and, in particular, the role of spatial context, both at the subcellular and tissue level, in shaping this molecular encoding.
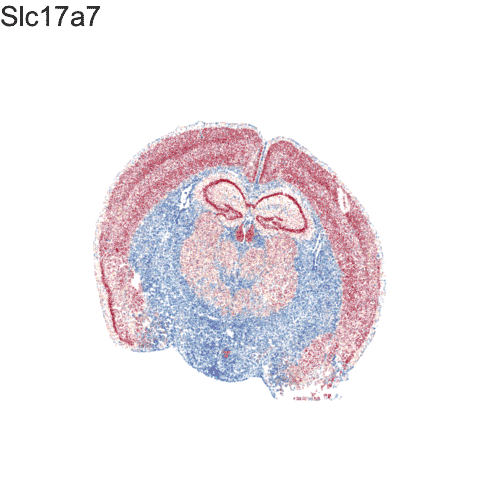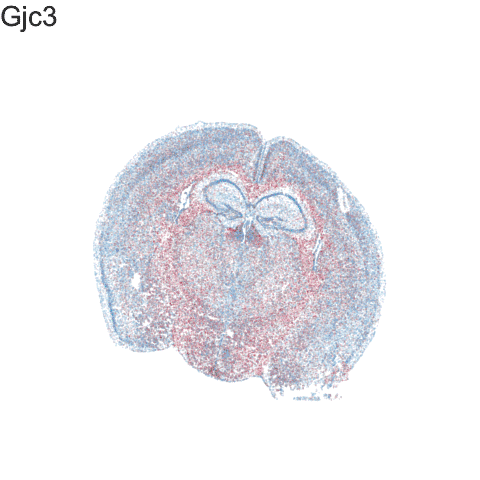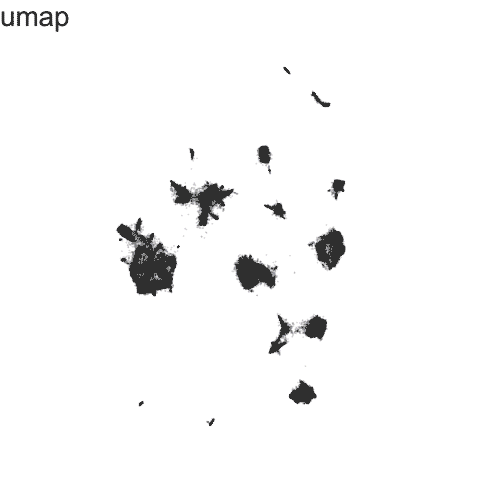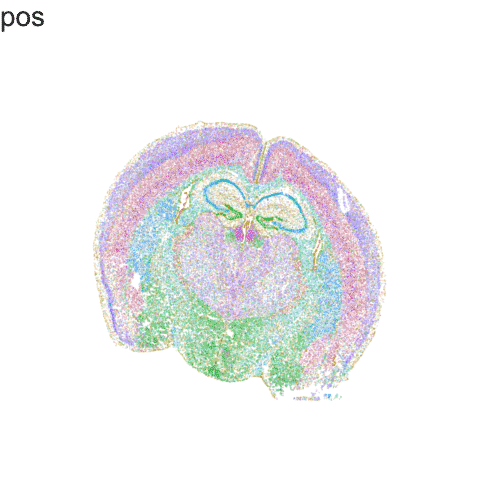
Recent technological advances have enabled spatially resolved molecular profiling to quantify gene and protein expression at molecule and single-cell resolution while preserving their spatial context within fixed cells and tissues in a high-throughput manner. Our lab specializes in the application of these technologies and the development of new machine learning and other statistical approaches as open-source computational software for analyzing such big single cell and spatially resolved omics datasets in order to take advantage of this new spatial information in deriving biological insights.
In this manner, we seek to better understand how a cell’s identity is established through the complex interplay of both intrinsic and extrinsic factors. We are interested in exploring these questions both in the context of healthy mammalian brain development as well as in the pathogenesis and progression of diseases such as pediatric gliomas, glioblastomas, and other brain cancers.